import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
# -----------------------------
# 1) Load data and set POT knobs
# -----------------------------
u = 2.5 # threshold [m]
dl_hours = 48 # declustering window [hours]
T = 100.0 # return period in years
df = pd.read_csv('Wavedata.csv') # adjust path if needed
df['datetime'] = pd.to_datetime(df['datetime'], dayfirst=True)
df['Hs'] = pd.to_numeric(df['Hs'], errors='coerce')
df = df.dropna(subset=['datetime', 'Hs']).sort_values('datetime').set_index('datetime')
# -------------------------------------------------------
# 2) Extract peaks over threshold with declustering (48 h)
# - keep only the local maximum within each cluster
# -------------------------------------------------------
exc = df.loc[df['Hs'] > u, 'Hs'] # raw exceedances
if exc.empty:
raise ValueError("No values exceed the threshold. Choose a lower u.")
# New cluster starts when the time gap since the previous exceedance > dl_hours
clusters = (exc.index.to_series().diff() > pd.Timedelta(hours=dl_hours)).cumsum()
# One peak per cluster: the maximum within that cluster
peaks = exc.groupby(clusters).max()
peak_times = exc.groupby(clusters).idxmax()
print(f'Number of exceedance clusters: {len(peaks)}')
print(f'First few peak times and values:\n{pd.Series(peaks.values, index=peak_times.values).head()}\n')
# -------------------------------------------------------
# 3) Fit GPD to exceedance magnitudes Y = X - u
# -------------------------------------------------------
y = (peaks - u).values
# SciPy's genpareto: c is xi, loc is usually fixed at 0 for exceedances
xi, loc, beta = stats.genpareto.fit(y, floc=0.0)
print('GPD fit parameters (exceedances):')
print(f' scale β = {beta:.4f}')
print(f' shape ξ = {xi:.4f}')
print(f' loc = {loc:.4f} (fixed at 0)\n')
# -------------------------------------------------------
# 4) 100-year return value using a Poisson rate of exceedances
# -------------------------------------------------------
# Estimate exceedance rate per year from the declustered peaks
years = (df.index.max() - df.index.min()) / pd.Timedelta(days=365.25)
rate = len(peaks) / float(years)
def gpd_return_level(T, u, beta, xi, rate, exact=True):
"""
Return level for POT with exceedance rate 'rate' per year.
exact=True uses the Poisson-exact formula.
"""
if exact:
# P(M <= z_T) = 1 - 1/T = exp(- rate * S(y_T)), with S(y) = 1 - G(y)
S = -np.log(1.0 - 1.0/T) / rate # survivor probability per exceedance
if np.isclose(xi, 0.0):
yT = -beta * np.log(S)
else:
yT = (beta/xi) * (S**(-xi) - 1.0)
else:
# Large-T approximation: S ≈ 1/(rate*T)
S = 1.0 / (rate * T)
if np.isclose(xi, 0.0):
yT = beta * np.log(1.0/S)
else:
yT = (beta/xi) * ((rate*T)**xi - 1.0)
return u + yT
z_T = gpd_return_level(T, u, beta, xi, rate, exact=True)
print(f'Estimated rate of exceedance: {rate:.3f} per year')
print(f'{int(T)}-year return value (exact Poisson formula): {z_T:.4f} m\n')
# Optional: finite upper end-point if xi < 0
if xi < 0:
x_max = u - beta/xi
print(f'Upper end-point of the model (xi < 0): {x_max:.4f} m\n')
# -------------------------------------------------------
# 5) Quick diagnostics: histogram+pdf and cdf for peaks
# -------------------------------------------------------
# Work in the original Hs scale for plotting
# Grid limited by support if xi < 0
y_max_support = np.inf if xi >= 0 else max(0.0, -beta/xi * 0.99)
y_max = y.max()
y_grid_max = min(y_max*1.5 if np.isfinite(y_max) else 1.0, y_max_support if np.isfinite(y_max_support) else y_max*1.5)
y_grid_max = max(y_grid_max, y.max()*1.2 + 1e-9)
yg = np.linspace(0, y_grid_max, 400)
xg = u + yg
pdf_g = stats.genpareto.pdf(yg, xi, loc=0, scale=beta)
cdf_g = stats.genpareto.cdf(yg, xi, loc=0, scale=beta)
# Empirical cdf for peaks in Hs
s = np.sort(peaks.values)
n = s.size
p_emp = np.arange(1, n+1) / (n + 1.0)
fig, axes = plt.subplots(1, 2, figsize=(11, 4.6), constrained_layout=True)
# Left: histogram of peaks in Hs with fitted GPD pdf on Hs scale
ax = axes[0]
ax.hist(peaks.values, bins='auto', density=True, facecolor='lightgrey',
edgecolor='indianred', alpha=0.7, label='Declustered peaks')
ax.plot(xg, pdf_g, color='red', lw=2,
label=fr'GPD over u={u} m $(\beta={beta:.2f},\, \xi={xi:.2f})$')
ax.axvline(u, color='grey', ls=':', lw=1.5, label='Threshold')
ax.axvline(z_T, color='crimson', ls='--', lw=2, label=fr'{int(T)}-yr RL = {z_T:.2f} m')
ax.set_xlabel('Hs (m)')
ax.set_ylabel('Density')
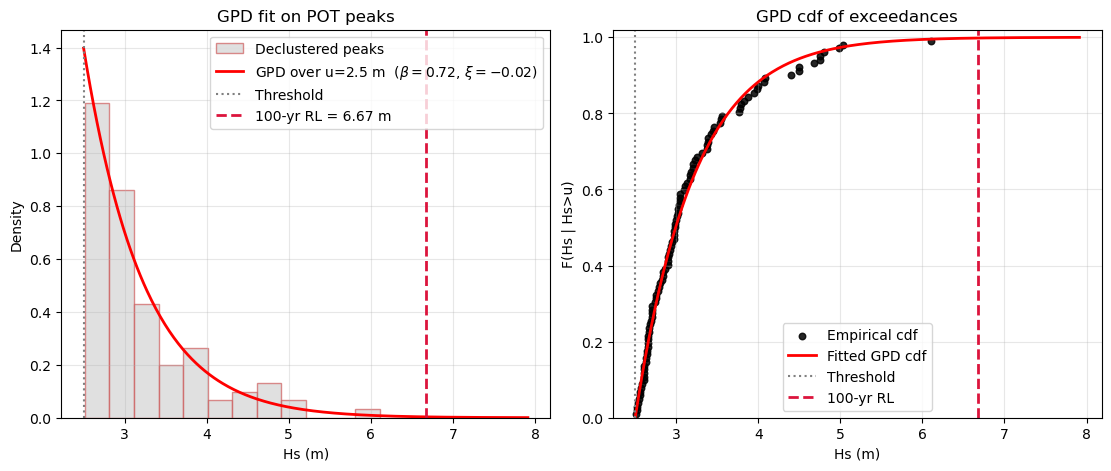
ax.set_title('GPD fit on POT peaks')
ax.legend(frameon=True)
ax.grid(alpha=0.3)
# Right: empirical cdf of peaks and fitted GPD cdf shown in Hs scale
ax = axes[1]
ax.scatter(s, p_emp, s=22, color='black', alpha=0.85, label='Empirical cdf')
ax.plot(xg, cdf_g, color='red', lw=2, label='Fitted GPD cdf')
ax.axvline(u, color='grey', ls=':', lw=1.5, label='Threshold')
ax.axvline(z_T, color='crimson', ls='--', lw=2, label=fr'{int(T)}-yr RL')
ax.set_xlabel('Hs (m)')
ax.set_ylabel('F(Hs | Hs>u)')
ax.set_ylim(0, 1.02)
ax.set_title('GPD cdf of exceedances')
ax.legend(frameon=True)
ax.grid(alpha=0.3)
plt.show()